|
The mean amino acid identities of ChdiNPV putative proteins with orthologous proteins of the above-mentioned 4 viruses were 98.3%, 87.7%, 87.4%, and 84.4%, respectively. A blastp search with a cutoff E value of 1 × 10 −5 using blast+ ( 9) found that ChdiNPV shared 140, 136, 136, and 136 ORFs with ChmuNPV, ChocNPV, CfMNPV, and ChroNPV, respectively. Global alignment using EMBOSS Stretcher ( 8) showed that the ChdiNPV genome had 95.4%, 81.6%, 81.2%, and 75.8% nucleotide identities to ChmuNPV, ChocNPV, CfMNPV, and ChroNPV, respectively. The remaining 144 ORFs had similarities to other baculovirus genes, including all baculovirus core genes. diversana NPV (ChdiNPV) was 122,827 bp long with a G+C content of 50.2% and was estimated to have 150 ORFs, including 6 ORFs unique to ChdiNPV. Putative open reading frames (ORFs) encoding more than 50 amino acids were identified using the Glimmer 3 version 1.5 ( 7) algorithm trained for ChmuNPV ORF annotations using Geneious Prime 2019.2.3 (Biomatters) and were manually edited. The genome sequence reported here used the sequence of the longer amplicon. The PCR generated two amplicons (482 and 656 bp) containing repeat sequences. An ambiguous region with lower coverage inside the orf1629 gene was amplified using PCR, and the DNA was sequenced using Sanger sequencing.  All processed reads were then used to assemble the circular contig, permitting 20,940,902 reads to be assembled and resulting in a mean coverage of 16,856× (standard deviation, 2,051×). This generated a contig with overlapping sequences at both ends, from which a circular contig was created. Assembly was conducted first using 5% of the total processed reads (1,052,696 of 21,053,934 reads) using de novo assembly in Geneious Prime 2019.2.3 (Biomatters) with default parameters of low sensitivity/fastest setting except that the “only use paired hits during assembly†option was used. Trimming of low-quality ends (Phred quality scores of  Pairwise comparisons between members of the genus Badnavirus showed that gooseberry vein banding associated virus GB1 (HQ852248) and rubus yellow net virus isolate Baumforth's Seedling A (KM078034) were the closest related virus sequences to SYLSV, sharing 73% identity at the nucleotide level. Both isolates have the same genome organization containing three open reading frames (ORFs), with ORF3 being the largest, encoding a putative polyprotein of 232 kDa with conserved domains including a zinc finger, pepsin-like aspartate protease, reverse transcriptase (RT), and RNase H. The sequences share 95% identity at the nucleotide level. 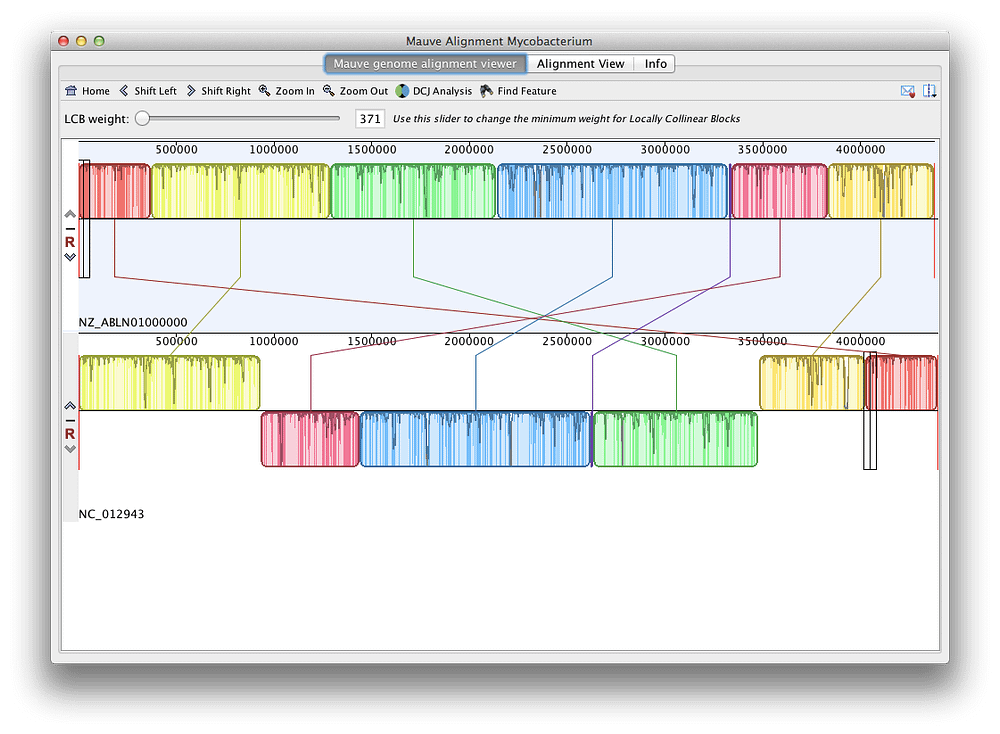 The viral genome of SYLSV-MN (Minnesota) and SYLSV-MD (Maryland) is 8,017bp in length. Spiraea ( Spiraea x bumalda) ‘Anthony Waterer’ plants showing virus-like symptoms including yellow spotting and leaf deformation were used for sequencing. 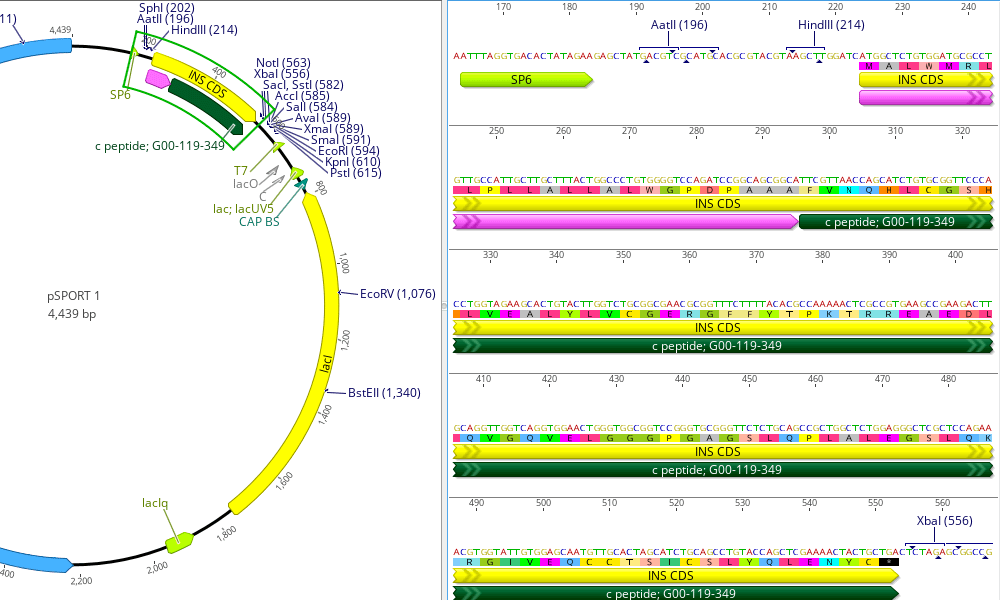 The complete genome sequences of two isolates of spiraea yellow leafspot virus (SYLSV) were determined.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
